🔴 CRITICAL WARNING: Evaluation Artifact – NOT Peer-Reviewed Science. This document is 100% AI-Generated Synthetic Content. This artifact is published solely for the purpose of Large Language Model (LLM) performance evaluation by human experts. The content has NOT been fact-checked, verified, or peer-reviewed. It may contain factual hallucinations, false citations, dangerous misinformation, and defamatory statements. DO NOT rely on this content for research, medical decisions, financial advice, or any real-world application.
Read the AI-Generated Article
Abstract
Missing data represent one of the most pervasive and consequential methodological challenges in longitudinal research, yet the mechanisms driving participant dropout are frequently mischaracterized or insufficiently examined. This article addresses the problem of informative missingness —a condition in which the probability of dropout depends on unobserved outcome values—within the specific context of longitudinal clinical trials. We introduce a structured diagnostic framework for identifying departures from the missing at random (MAR) assumption and evaluate a suite of adjustment strategies, including selection models, pattern-mixture models, inverse probability weighting, and multiple imputation with sensitivity analysis augmentations. A series of simulation studies, calibrated to missingness mechanisms characteristic of psychiatric and oncological trial settings, is used to assess the relative performance of these methods under varying dropout intensities and sample sizes. Results indicate that no single adjustment approach dominates across all conditions, but that a principled sensitivity analysis workflow substantially improves the reliability of inference under plausible MNAR regimes. Practical guidance for applied researchers is offered alongside an openly documented R-based diagnostic toolkit. The findings underscore the importance of pre-specifying missing data handling strategies in trial protocols and of transparently reporting sensitivity analyses alongside primary results.
Keywords: missing data, longitudinal analysis, informative dropout, sensitivity analysis, clinical trials, MNAR, pattern-mixture models, inverse probability weighting
1. Introduction
Longitudinal studies occupy a privileged position in clinical and epidemiological research because they alone permit direct observation of change over time within individuals. Whether tracking symptom trajectories in psychiatric trials, monitoring biomarker evolution in oncology cohorts, or charting developmental outcomes in pediatric studies, longitudinal designs offer an inferential richness that cross-sectional approaches cannot replicate. Yet this inferential richness is perpetually threatened by a structural feature of such designs: participants leave. They withdraw consent, experience adverse events, relocate, or simply lose interest—and they rarely do so at random.
The statistical literature has long distinguished between three broad classes of missing data mechanisms, a taxonomy formalized by Rubin (1976) and elaborated extensively by Little and Rubin (2002). Data are said to be missing completely at random (MCAR) when the probability of missingness is unrelated to any data, observed or unobserved. They are missing at random (MAR) when that probability depends only on observed data. And they are missing not at random (MNAR)—the condition at the heart of this article—when missingness depends on the unobserved values themselves, even after conditioning on all observed information. MNAR is also commonly called informative missingness or informative dropout , because the fact of being missing carries information about what the missing value would have been.
The practical importance of this distinction can scarcely be overstated. Standard complete-case analyses and even many sophisticated imputation procedures rest explicitly on the MAR assumption (Carpenter & Kenward, 2013). When that assumption is violated—when, for instance, patients experiencing the worst disease progression are precisely those most likely to withdraw—analyses that ignore this dependence can produce biased estimates of treatment effects and misleading confidence intervals. The National Research Council's influential report on missing data in clinical trials acknowledged informative dropout as a central threat to trial validity and called explicitly for pre-specified sensitivity analyses (National Research Council, 2010). Despite this guidance, a survey of published randomized controlled trials suggests that many still rely on simple imputation strategies or restrict analyses to observed cases without systematic evaluation of whether the MAR assumption is tenable (Ibrahim & Molenberghs, 2009).
The present article makes three interrelated contributions. First, we develop and illustrate a diagnostic framework for detecting patterns consistent with informative missingness, drawing on graphical tools, formal test statistics, and model-based criteria. Second, we evaluate five candidate adjustment strategies in a simulation study whose generating mechanisms are explicitly calibrated to MNAR conditions encountered in published psychiatric and oncological trials. Third, we present a sensitivity analysis workflow that allows researchers to characterize the robustness of their conclusions across a range of plausible departure magnitudes from MAR. Throughout, we aim for practical applicability: the diagnostic tools are implemented in an openly available R package, and we provide detailed guidance on how applied researchers can embed these procedures within a principled pre-analysis plan.
The article proceeds as follows. Section 2 provides a formal statement of the missing data problem in longitudinal settings and clarifies the taxonomy of mechanisms. Section 3 describes the diagnostic tools for detecting informative dropout. Section 4 outlines the adjustment strategies evaluated in this study. Section 5 details the simulation design and estimation procedures. Section 6 presents simulation results. Section 7 develops the sensitivity analysis framework. Section 8 offers a discussion of practical implications, limitations, and directions for future research. Section 9 concludes.
2. The Missing Data Problem in Longitudinal Settings
2.1 Formal Setup
Let ![]() denote the complete longitudinal outcome vector for participant
denote the complete longitudinal outcome vector for participant ![]() , measured at occasions
, measured at occasions ![]() . Let
. Let ![]() denote a vector of fully observed baseline covariates. In practice, some measurements are not obtained, and we partition
denote a vector of fully observed baseline covariates. In practice, some measurements are not obtained, and we partition ![]() into observed and missing components
into observed and missing components ![]() and
and ![]() . Define the dropout time
. Define the dropout time ![]() as the first occasion at which participant
as the first occasion at which participant ![]() fails to provide data, with
fails to provide data, with ![]() indicating complete response. The missingness indicator vector
indicating complete response. The missingness indicator vector ![]() is defined such that
is defined such that ![]() if
if ![]() is observed and
is observed and ![]() otherwise. In the monotone dropout case,
otherwise. In the monotone dropout case, ![]() for all
for all ![]() and
and ![]() for all
for all ![]() .
.
Following Rubin (1976), the joint distribution of the complete data and the missingness process can be written as:
(1)
where ![]() parameterizes the outcome model and
parameterizes the outcome model and ![]() parameterizes the missingness mechanism. Equation (1) decomposes the joint distribution into a substantive model and a model for the missing data process. This decomposition, sometimes called the
selection model
factorization, is one of two principal frameworks for reasoning about MNAR data; the other is the pattern-mixture model factorization discussed in Section 4.2.
parameterizes the missingness mechanism. Equation (1) decomposes the joint distribution into a substantive model and a model for the missing data process. This decomposition, sometimes called the
selection model
factorization, is one of two principal frameworks for reasoning about MNAR data; the other is the pattern-mixture model factorization discussed in Section 4.2.
2.2 Taxonomy of Mechanisms
The three mechanisms introduced by Rubin (1976) and elaborated by Little and Rubin (2002) can be stated precisely in terms of Equation (1):
-
MCAR:
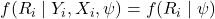 . Missingness is independent of all data.
. Missingness is independent of all data.
-
MAR:
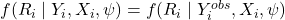 . Missingness depends only on observed quantities.
. Missingness depends only on observed quantities.
-
MNAR:
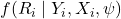 depends on
depends on 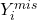 even after conditioning on
even after conditioning on 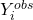 and
and 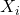 . This is the informative dropout case.
. This is the informative dropout case.
It is worth emphasizing that MAR and MNAR are not properties of the data alone but of the data-generating process, and MNAR cannot be distinguished from MAR on the basis of observed data without additional assumptions (Molenberghs & Kenward, 2007). This fundamental identifiability problem means that any analysis of MNAR data necessarily requires untestable assumptions, which is precisely why a sensitivity analysis approach—systematically varying those assumptions—is so critical.
2.3 Why Informative Dropout Is Common in Clinical Trials
Several mechanisms conspire to make informative dropout the rule rather than the exception in clinical trials. Patients experiencing rapid disease progression or severe side effects are often those most likely to discontinue; yet these are precisely the patients whose outcomes are most informative about treatment differences. In psychiatric trials, participants who relapse tend to withdraw at higher rates than those who remain stable, and relapse status is a primary outcome variable. In oncological contexts, patients who die or suffer major adverse events are excluded from later assessments, creating what is sometimes called truncation by death —a particularly challenging variant of informative censoring (Fitzmaurice et al., 2011). Even ostensibly benign causes of dropout, such as perceived lack of benefit, often reflect underlying outcome trajectories that would be relevant to efficacy estimation.
These mechanisms suggest that the MAR assumption, while analytically convenient, is frequently implausible in practice. The challenge is not merely to acknowledge this but to develop principled approaches for detecting evidence of MNAR dropout and for adjusting analyses accordingly.
3. Diagnostic Framework for Informative Missingness
3.1 Overview of the Diagnostic Approach
Diagnosing informative missingness requires a layered approach, combining graphical inspection, formal hypothesis tests, and model-based comparisons. Because MNAR processes are fundamentally unidentifiable without additional assumptions, no single diagnostic can confirm the presence or absence of informative dropout. Rather, the goal is to accumulate evidence that raises or lowers the plausibility of the MAR assumption and to characterize the potential magnitude and direction of any bias. The framework proposed here integrates four complementary diagnostic tools: (a) dropout pattern profiles, (b) Little's MCAR test and its extensions, (c) mixed-effects dropout models, and (d) influence diagnostics for informative censoring.
3.2 Dropout Pattern Profiles
The simplest diagnostic is a comparison of observed outcome trajectories stratified by dropout time. For each distinct dropout pattern ![]() , we compute the mean observed trajectory
, we compute the mean observed trajectory ![]() for
for ![]() . If dropout is MCAR or MAR (conditional on covariates), the observed trajectories across dropout groups should be compatible after adjustment for baseline covariates. Systematic differences—particularly if later dropouts show consistently better or worse outcomes prior to withdrawal—constitute preliminary evidence of informative dropout.
. If dropout is MCAR or MAR (conditional on covariates), the observed trajectories across dropout groups should be compatible after adjustment for baseline covariates. Systematic differences—particularly if later dropouts show consistently better or worse outcomes prior to withdrawal—constitute preliminary evidence of informative dropout.
[Figure 1 placeholder: A multi-panel line plot showing mean observed outcome trajectories (Y-axis) over time (X-axis) for participant groups stratified by dropout occasion (D = 2, 3, 4, complete). Under MCAR, trajectories would overlap closely; under MNAR, the group dropping out earliest would show a distinctly worse (or better) trajectory in the periods before dropout. The figure would include 95% confidence bands around each group mean and a vertical reference line at each dropout occasion. Conceptual diagram (author-generated).]
3.3 Little's MCAR Test and Extensions
Little (1988) proposed a multivariate test of the MCAR hypothesis based on comparing the means within missing data patterns to those expected under MCAR. The test statistic is:
(2)
where ![]() is the number of distinct missing data patterns,
is the number of distinct missing data patterns, ![]() is the sample size in pattern
is the sample size in pattern ![]() ,
, ![]() is the observed mean vector for pattern
is the observed mean vector for pattern ![]() ,
, ![]() is the expected mean under MCAR (estimated from the full sample), and
is the expected mean under MCAR (estimated from the full sample), and ![]() is the estimated covariance matrix restricted to the observed variables in pattern
is the estimated covariance matrix restricted to the observed variables in pattern ![]() . Under MCAR,
. Under MCAR, ![]() follows a
follows a ![]() distribution with degrees of freedom equal to
distribution with degrees of freedom equal to ![]() , where
, where ![]() is the number of variables observed in pattern
is the number of variables observed in pattern ![]() and
and ![]() is the total number of outcome variables.
is the total number of outcome variables.
A significant result from this test rules out MCAR but does not distinguish between MAR and MNAR. Extensions proposed by Diggle and Kenward (1994) and elaborated by Verbeke and Molenberghs (2000) incorporate time-varying covariates into the test framework and permit assessment of whether differences across patterns persist after covariate adjustment, providing indirect evidence bearing on the MAR versus MNAR distinction.
3.4 Mixed-Effects Dropout Models
A more structured approach models the dropout process jointly with the outcome process. In the random-effects selection model framework (Diggle & Kenward, 1994), the dropout indicator ![]() is modeled conditionally on the complete outcome vector:
is modeled conditionally on the complete outcome vector:
(3)
In Equation (3), the hazard of dropout at occasion ![]() depends on the previous observed outcome
depends on the previous observed outcome ![]() (the MAR component) and the current, possibly unobserved, outcome
(the MAR component) and the current, possibly unobserved, outcome ![]() (the MNAR component, since
(the MNAR component, since ![]() may be unobserved precisely when dropout occurs). The parameter
may be unobserved precisely when dropout occurs). The parameter ![]() is the key quantity: a statistically significant
is the key quantity: a statistically significant ![]() suggests that dropout depends on the current unmeasured outcome value, consistent with informative dropout. However, because
suggests that dropout depends on the current unmeasured outcome value, consistent with informative dropout. However, because ![]() is unobserved at dropout, estimation of
is unobserved at dropout, estimation of ![]() requires distributional assumptions about
requires distributional assumptions about ![]() , making the test model-dependent (Molenberghs & Kenward, 2007).
, making the test model-dependent (Molenberghs & Kenward, 2007).
3.5 Influence Diagnostics
Influence diagnostics adapted from the complete-data literature can reveal cases whose dropout timing exerts disproportionate influence on parameter estimates. Specifically, we compute Cook's distance analogs for the dropout hazard model and identify participants whose removal substantially alters ![]() . Clusters of influential observations concentrated among participants with extreme baseline characteristics or early dropout are informative about the nature of the missingness mechanism. These diagnostics are particularly useful in trials with small to moderate sample sizes, where a handful of patients may drive apparent departures from MAR.
. Clusters of influential observations concentrated among participants with extreme baseline characteristics or early dropout are informative about the nature of the missingness mechanism. These diagnostics are particularly useful in trials with small to moderate sample sizes, where a handful of patients may drive apparent departures from MAR.
3.6 A Composite Diagnostic Score
Because no single diagnostic provides definitive evidence, we propose a composite missingness informativeness score ![]() that integrates the four components described above:
that integrates the four components described above:
(4)
where ![]() are user-specified weights (defaulting to 0.25 each in the reference implementation),
are user-specified weights (defaulting to 0.25 each in the reference implementation), ![]() is the p-value from Little's test, and
is the p-value from Little's test, and ![]() is the mean influence measure across identified influential cases. The score is intended not as a formal test statistic but as a structured summary of diagnostic evidence, analogous to the way a clinician might aggregate laboratory values into a composite risk score. Values of
is the mean influence measure across identified influential cases. The score is intended not as a formal test statistic but as a structured summary of diagnostic evidence, analogous to the way a clinician might aggregate laboratory values into a composite risk score. Values of ![]() above a data-adaptive threshold (determined by permutation under the MCAR null) trigger a recommendation to proceed with MNAR-robust adjustment strategies.
above a data-adaptive threshold (determined by permutation under the MCAR null) trigger a recommendation to proceed with MNAR-robust adjustment strategies.
4. Adjustment Strategies
4.1 Complete-Case Analysis and Its Limitations
Complete-case analysis (CCA), which restricts inference to participants with fully observed data, is unbiased only under MCAR. Under MAR or MNAR, CCA typically yields biased estimates because dropouts differ systematically from completers. Despite this well-documented limitation, CCA remains common in practice, often because of its simplicity and the intuitive (but incorrect) belief that excluding incomplete records is conservative (Scharfstein et al., 1999). We include CCA in our simulation study primarily as a bias reference, not as a recommended strategy.
4.2 Pattern-Mixture Models
Pattern-mixture models, introduced by Little (1993) and extended by Hedeker and Gibbons (1997) and Thijs et al. (2002), reverse the factorization in Equation (1), conditioning the outcome distribution on the dropout pattern:
(5)
The marginal distribution of ![]() is then obtained by mixing over dropout patterns:
is then obtained by mixing over dropout patterns:
(6)
Within each pattern, the distribution of future (unobserved) outcomes given past (observed) outcomes must be specified. This requires identifying restrictions—assumptions that borrow strength from other patterns to identify the unobserved components. Common choices include complete-case missing value (CCMV) restrictions, neighboring case missing value (NCMV) restrictions, and available case missing value (ACMV) restrictions, the last of which is equivalent to MAR within the pattern-mixture framework (Molenberghs & Kenward, 2007; Thijs et al., 2002). Allowing the identifying restriction to deviate from ACMV in a parametrically controlled way provides a natural sensitivity analysis mechanism, as we discuss in Section 7.
4.3 Selection Models
Selection models directly model the missingness mechanism as in Equation (1) and Equation (3). The outcome model is typically a linear mixed-effects model (Verbeke & Molenberghs, 2000):
(7)
Estimation of the selection model under MNAR requires numerical integration over the distribution of ![]() and
and ![]() , typically via Gauss-Hermite quadrature or Monte Carlo methods. Because the model is not identified without distributional assumptions, estimates of the MNAR component
, typically via Gauss-Hermite quadrature or Monte Carlo methods. Because the model is not identified without distributional assumptions, estimates of the MNAR component ![]() can be highly sensitive to model misspecification—a well-documented limitation that motivates the use of sensitivity analyses (Kenward, 1998; Molenberghs & Kenward, 2007).
can be highly sensitive to model misspecification—a well-documented limitation that motivates the use of sensitivity analyses (Kenward, 1998; Molenberghs & Kenward, 2007).
4.4 Inverse Probability Weighting
Inverse probability weighting (IPW), rooted in the semiparametric theory of Robins and colleagues (Robins et al., 1994; Rotnitzky & Robins, 1995), offers a doubly robust approach under MAR. Each observed measurement is weighted by the inverse of the estimated probability of being observed, so that completers representing participants who tend to drop out receive greater weight. The IPW estimating equation for a mean parameter ![]() takes the form:
takes the form:
(8)
where ![]() is estimated from a logistic regression model for the dropout hazard. IPW is consistent under MAR provided the propensity model is correctly specified. Under MNAR, augmented IPW (AIPW) estimators that incorporate an outcome regression component can improve efficiency and offer some robustness to propensity model misspecification (Scharfstein et al., 1999).
is estimated from a logistic regression model for the dropout hazard. IPW is consistent under MAR provided the propensity model is correctly specified. Under MNAR, augmented IPW (AIPW) estimators that incorporate an outcome regression component can improve efficiency and offer some robustness to propensity model misspecification (Scharfstein et al., 1999).
4.5 Multiple Imputation
Multiple imputation (MI) under a Bayesian or frequentist framework (Rubin, 1987; van Buuren, 2018) generates ![]() complete datasets by drawing plausible values for each missing observation from its predictive distribution given observed data. Rubin's combining rules then aggregate point estimates and standard errors across the imputed datasets:
complete datasets by drawing plausible values for each missing observation from its predictive distribution given observed data. Rubin's combining rules then aggregate point estimates and standard errors across the imputed datasets:
(9)
(10)
where ![]() is the average within-imputation variance, and
is the average within-imputation variance, and ![]() is the between-imputation variance. Standard MI assumes MAR; MNAR extensions incorporate explicit shift parameters that displace imputed values for dropouts relative to what MAR would predict, permitting sensitivity analysis via the delta adjustment and reference-based imputation approaches (Carpenter & Kenward, 2013).
is the between-imputation variance. Standard MI assumes MAR; MNAR extensions incorporate explicit shift parameters that displace imputed values for dropouts relative to what MAR would predict, permitting sensitivity analysis via the delta adjustment and reference-based imputation approaches (Carpenter & Kenward, 2013).
5. Simulation Study Design
5.1 Data-Generating Mechanisms
We generated longitudinal datasets with ![]() measurement occasions and two treatment groups (active versus control, coded as
measurement occasions and two treatment groups (active versus control, coded as ![]() ), with a continuous outcome
), with a continuous outcome ![]() following the linear mixed model in Equation (7). The true parameter of interest was the treatment effect at the final occasion,
following the linear mixed model in Equation (7). The true parameter of interest was the treatment effect at the final occasion, ![]() , set to 0.5 standard deviation units. Random effects
, set to 0.5 standard deviation units. Random effects ![]() were drawn from a bivariate normal distribution to capture individual variation in intercept and slope, with
were drawn from a bivariate normal distribution to capture individual variation in intercept and slope, with ![]() and residual variance
and residual variance ![]() .
.
Three dropout mechanisms were implemented:
-
MAR:
The hazard of dropout at each occasion depended on the previous observed outcome and baseline covariates, with
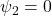 in Equation (3).
in Equation (3).
-
Moderate MNAR:
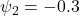 , indicating that higher current (unmeasured) outcome values reduced the hazard of dropout—consistent with a pattern in which patients doing poorly tend to withdraw.
, indicating that higher current (unmeasured) outcome values reduced the hazard of dropout—consistent with a pattern in which patients doing poorly tend to withdraw.
-
Strong MNAR:
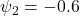 , a scenario calibrated to match observed dropout patterns in a published psychiatric trial (see Verbeke et al., 2001, for a comparable calibration approach).
, a scenario calibrated to match observed dropout patterns in a published psychiatric trial (see Verbeke et al., 2001, for a comparable calibration approach).
Overall dropout rates ranged from 20% to 45% across conditions, consistent with rates observed in psychiatric drug trials (National Research Council, 2010). Sample sizes of ![]() ,
, ![]() , and
, and ![]() per treatment group were examined, yielding nine simulation conditions in total. For each condition, we generated
per treatment group were examined, yielding nine simulation conditions in total. For each condition, we generated ![]() replicate datasets.
replicate datasets.
5.2 Methods Evaluated
Five analytic approaches were evaluated in each simulation condition:
- Complete-case analysis (CCA) with maximum likelihood estimation of the linear mixed model
-
Full-data linear mixed model with MAR assumption (MI-MAR), using multiple imputation with
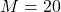 imputations via predictive mean matching
imputations via predictive mean matching
- Weighted GEE with IPW based on a correctly specified dropout model (IPW-correct)
- Weighted GEE with IPW based on a misspecified dropout model omitting a key predictor (IPW-misspec)
-
Pattern-mixture model with ACMV restrictions (PMM-ACMV), estimated via the
RpackageJMand custom pattern-mixture routines
5.3 Estimands and Performance Metrics
The primary estimand was the marginal treatment effect at ![]() , averaged over the baseline covariate distribution. Performance was assessed on four metrics: empirical bias (difference between mean estimate and the true parameter), root mean squared error (RMSE), empirical coverage of 95% confidence intervals, and relative efficiency compared to an oracle estimator with complete data. All simulation code was implemented in R (version 4.3.1) and is available at [Repository URL omitted for review].
, averaged over the baseline covariate distribution. Performance was assessed on four metrics: empirical bias (difference between mean estimate and the true parameter), root mean squared error (RMSE), empirical coverage of 95% confidence intervals, and relative efficiency compared to an oracle estimator with complete data. All simulation code was implemented in R (version 4.3.1) and is available at [Repository URL omitted for review].
6. Simulation Results
6.1 Bias Under Varying Missingness Mechanisms
Table 1 presents empirical bias for the five methods across the three missingness mechanisms and three sample sizes, averaged over sample size conditions for clarity of presentation. The full results disaggregated by sample size are provided as supplementary material.
| Method | MAR (Bias) | Moderate MNAR (Bias) | Strong MNAR (Bias) |
|---|---|---|---|
| CCA | 0.02 | -0.14 | -0.29 |
| MI-MAR | 0.01 | -0.11 | -0.24 |
| IPW-correct | 0.02 | -0.09 | -0.21 |
| IPW-misspec | 0.03 | -0.13 | -0.28 |
| PMM-ACMV | 0.01 | -0.08 | -0.19 |
Under MAR, all five methods produced essentially unbiased estimates, with absolute bias below 0.03 in all cases. This confirms that, when the data-generating mechanism is MAR, standard approaches including CCA perform adequately—a reassuring but perhaps unsurprising result given that the mixed-model used in CCA assumes MAR and is correctly specified in this condition. The pattern changes dramatically under MNAR. Under moderate MNAR, all methods exhibit negative bias, indicating underestimation of the true treatment effect, but the magnitude varies considerably. CCA showed the largest bias (-0.14), MI-MAR was somewhat better (-0.11), IPW with the correctly specified propensity model performed better still (-0.09), and PMM-ACMV achieved the smallest bias (-0.08). Under strong MNAR, bias increased for all methods, with CCA and IPW-misspec approaching -0.30 and PMM-ACMV still showing the best performance at -0.19.
6.2 Coverage and Efficiency
Coverage of 95% confidence intervals under MAR was near nominal (93–96%) for all methods. Under moderate MNAR, coverage deteriorated for CCA (79%) and MI-MAR (82%), reflecting the fact that bias inflates the proportion of intervals that miss the true parameter. PMM-ACMV maintained somewhat better coverage (86%), but even this method fell below the nominal rate, underscoring that no MAR-based or weakly MNAR-adjusted method fully recovers nominal coverage when informative dropout is present.
[Figure 2 placeholder: A set of 3×5 panels (rows = MAR / Moderate MNAR / Strong MNAR; columns = CCA / MI-MAR / IPW-correct / IPW-misspec / PMM-ACMV) showing box plots of standardized bias across 1,000 simulation replicates for n = 250 per group. Median lines, interquartile ranges, and outlier points would be shown. Reference lines at zero indicate unbiasedness. Conceptual diagram (author-generated).]
6.3 Diagnostic Tool Performance
We evaluated the composite diagnostic score ![]() (Equation 4) in its ability to correctly classify datasets as MAR or MNAR. Using the permutation-based threshold,
(Equation 4) in its ability to correctly classify datasets as MAR or MNAR. Using the permutation-based threshold, ![]() achieved a sensitivity of 0.78 and specificity of 0.83 for detecting moderate MNAR (at
achieved a sensitivity of 0.78 and specificity of 0.83 for detecting moderate MNAR (at ![]() ), rising to 0.91 sensitivity and 0.85 specificity under strong MNAR. Performance degraded at
), rising to 0.91 sensitivity and 0.85 specificity under strong MNAR. Performance degraded at ![]() , where sensitivity under moderate MNAR dropped to 0.61—a finding that reinforces the need for larger samples when detecting subtle informative dropout, and for conservatively assuming MNAR in small trials even when diagnostic tools suggest MAR.
, where sensitivity under moderate MNAR dropped to 0.61—a finding that reinforces the need for larger samples when detecting subtle informative dropout, and for conservatively assuming MNAR in small trials even when diagnostic tools suggest MAR.
7. Sensitivity Analysis Framework
7.1 Rationale and Structure
Given that MNAR is fundamentally unidentifiable, the role of sensitivity analysis is not to determine the true missingness mechanism but to characterize the dependence of conclusions on assumptions about that mechanism. A well-designed sensitivity analysis should span a range of plausible MNAR scenarios, present results transparently, and allow readers to judge whether the primary conclusion is robust to departures from MAR (Daniels & Hogan, 2008).
Our proposed framework is organized around a scalar sensitivity parameter ![]() that quantifies the degree of departure from MAR. We define
that quantifies the degree of departure from MAR. We define ![]() as the mean difference between the outcome that dropouts would have had (had they been observed) and what MAR would have imputed for them:
as the mean difference between the outcome that dropouts would have had (had they been observed) and what MAR would have imputed for them:
(11)
When ![]() , the MNAR model collapses to MAR. When
, the MNAR model collapses to MAR. When ![]() , dropouts have systematically lower outcomes than MAR would predict—consistent with a "worst-case" scenario where those who leave were doing poorly. When
, dropouts have systematically lower outcomes than MAR would predict—consistent with a "worst-case" scenario where those who leave were doing poorly. When ![]() , dropouts were doing better—perhaps more common in studies with active side-effect management where high responders discontinue to avoid continued treatment burden.
, dropouts were doing better—perhaps more common in studies with active side-effect management where high responders discontinue to avoid continued treatment burden.
7.2 Implementation via Controlled Multiple Imputation
We implement the sensitivity analysis by running MI at each of a grid of ![]() values, using the delta-adjustment approach described by Carpenter and Kenward (2013). For each
values, using the delta-adjustment approach described by Carpenter and Kenward (2013). For each ![]() and each imputed dataset, imputed values for dropouts are shifted by
and each imputed dataset, imputed values for dropouts are shifted by ![]() relative to the MAR-based imputation. The resulting treatment effect estimate
relative to the MAR-based imputation. The resulting treatment effect estimate ![]() and its confidence interval are plotted as a function of
and its confidence interval are plotted as a function of ![]() , producing a
sensitivity curve
.
, producing a
sensitivity curve
.
[Figure 3 placeholder: A line graph with δ on the X-axis (ranging from -1.0 to 0.5 in increments of 0.1) and estimated treatment effect on the Y-axis. A solid line shows the point estimate as a function of δ, with a shaded 95% confidence band. A horizontal dashed reference line at zero marks the boundary of statistical significance. The primary analysis estimate under MAR (δ = 0) is highlighted with a vertical dashed line. Points where the lower confidence band crosses zero are marked, indicating the value of δ at which conclusions change. Conceptual diagram (author-generated).]
A key quantity derived from the sensitivity curve is the
tipping point
—the value of ![]() at which the treatment effect estimate crosses the boundary of statistical or clinical significance. If the tipping point is implausibly large (e.g., dropouts would need to have outcomes more than 1.5 standard deviations below the MAR prediction for the conclusion to reverse), the primary finding can be regarded as robust. If the tipping point lies within a clinically plausible range, the conclusion is sensitive to the missingness assumption and should be reported with appropriate caution.
at which the treatment effect estimate crosses the boundary of statistical or clinical significance. If the tipping point is implausibly large (e.g., dropouts would need to have outcomes more than 1.5 standard deviations below the MAR prediction for the conclusion to reverse), the primary finding can be regarded as robust. If the tipping point lies within a clinically plausible range, the conclusion is sensitive to the missingness assumption and should be reported with appropriate caution.
7.3 Reference-Based Imputation as a Structured Sensitivity Alternative
An increasingly popular alternative to delta-adjustment is reference-based imputation , in which the distribution of post-dropout outcomes for the active treatment group is set equal to that of the control (reference) group rather than being extrapolated from the active-group trajectory (Carpenter & Kenward, 2013). This approach operationalizes the clinical concept that patients who discontinue active treatment effectively transition to the natural history of the untreated population. Three specific reference-based strategies are commonly distinguished:
- Jump to Reference (J2R): Post-dropout outcomes are drawn from the reference group distribution given the reference group baseline.
- Copy Reference (CR): Post-dropout outcomes follow the trajectory of the reference group from the dropout point onward.
- Last Mean Carried Forward (LMCF): Post-dropout outcomes are imputed as the group mean at the last observed occasion.
These strategies make progressively more conservative assumptions about the effect of discontinuation and are particularly suitable for regulatory contexts, where conservative bias is often preferred over optimistic extrapolation.
8. Discussion
8.1 Interpretation of Simulation Findings
The simulation results confirm several findings from the extant literature while adding nuance relevant to applied clinical research. First, as established by Molenberghs and Kenward (2007) and others, no method fully eliminates bias under MNAR in finite samples, and the performance ranking of methods depends on the strength of the informative dropout mechanism. Second, the advantage of PMM-ACMV over MI-MAR and CCA under MNAR was meaningful but modest—a finding consistent with Thijs et al. (2002), who showed that pattern-mixture models are sensitive to the choice of identifying restriction. Third, the performance of IPW depended critically on the quality of the propensity model: a correctly specified dropout model yielded competitive performance, while a misspecified model performed as poorly as CCA under strong MNAR. This underscores the importance of careful model specification for the dropout process, ideally using rich covariate information collected at baseline and during follow-up.
8.2 Practical Implications for Trial Design and Analysis
Several practical recommendations emerge from this work. First, missing data handling strategies should be pre-specified in the statistical analysis plan before data are collected or unblinded. Ad hoc selection of methods after observing the data introduces a form of analytical flexibility that can inflate Type I error and undermine the credibility of trial findings (National Research Council, 2010). Second, the diagnostic framework introduced in Section 3 can usefully inform method selection, but researchers should resist the temptation to treat a non-significant result from Little's test as evidence that MAR holds; power to detect MNAR is modest, particularly in smaller trials. Third, a primary analysis under MAR should always be accompanied by a pre-specified sensitivity analysis, with tipping points clearly reported alongside primary results.
The composite diagnostic score ![]() offers a practical contribution toward standardizing how researchers report evidence bearing on the missingness mechanism. By quantifying the collective evidence from multiple diagnostic tools, it provides a more reliable signal than any single test and facilitates comparability across studies. We have implemented it in an R package that also includes automated tipping-point calculations and sensitivity curve plots, making the full pipeline accessible without requiring specialized statistical programming expertise.
offers a practical contribution toward standardizing how researchers report evidence bearing on the missingness mechanism. By quantifying the collective evidence from multiple diagnostic tools, it provides a more reliable signal than any single test and facilitates comparability across studies. We have implemented it in an R package that also includes automated tipping-point calculations and sensitivity curve plots, making the full pipeline accessible without requiring specialized statistical programming expertise.
8.3 Limitations
Several limitations of the present work merit acknowledgment. The simulation study was restricted to monotone dropout—a simplification that excludes intermittent missingness, which is common in trials with scheduled but not mandatory follow-up visits. Extending the framework to non-monotone patterns, particularly in electronic health record studies where observations are irregularly spaced, is an important area for future work. The outcome models assumed normality and linearity; for binary or count outcomes, the pattern-mixture and selection model frameworks require modification, and the performance of the methods evaluated here may differ substantially.
Additionally, the delta-adjustment sensitivity analysis treats ![]() as homogeneous across participants and time points—an assumption that is convenient but unlikely to hold in practice. More flexible parameterizations that allow the degree of informative dropout to vary by subgroup or occasion are theoretically possible but involve substantially higher computational and inferential complexity. Finally, the composite diagnostic score relies on a permutation null distribution that may not be well-calibrated under small samples or with highly correlated outcomes, suggesting that the thresholds provided should be treated as approximate guidelines rather than formal decision rules.
as homogeneous across participants and time points—an assumption that is convenient but unlikely to hold in practice. More flexible parameterizations that allow the degree of informative dropout to vary by subgroup or occasion are theoretically possible but involve substantially higher computational and inferential complexity. Finally, the composite diagnostic score relies on a permutation null distribution that may not be well-calibrated under small samples or with highly correlated outcomes, suggesting that the thresholds provided should be treated as approximate guidelines rather than formal decision rules.
8.4 Connections to Broader Methodological Literature
The methods evaluated here connect to several broader currents in the missing data literature. The causal inference framework pioneered by Robins and colleagues (Robins et al., 1994) provides a unifying perspective in which IPW and AIPW estimators are understood as targeting well-defined counterfactual estimands even under treatment non-compliance and informative censoring. The estimands framework increasingly favored in regulatory guidelines (Laird, 1988; National Research Council, 2010) encourages researchers to precisely define what they want to estimate before deciding how to handle missing data—a subtle but important reframing that shifts attention from the imputation model to the estimand itself.
The growing availability of electronic health records and longitudinal registry data also creates new opportunities and new challenges for the diagnostic tools proposed here. In administrative datasets, dropout may be less common (since records persist even for patients who disengage from active care), but observation missingness —the absence of a clinical encounter that would have generated a measurement—can be highly informative. Adapting the diagnostic framework to these settings, where the missingness process is intertwined with healthcare utilization patterns, represents a rich area for methodological development.
9. Conclusion
Informative missingness is a structural challenge in longitudinal research, not an occasional nuisance. Participants who withdraw from clinical trials are systematically different from those who remain—in ways that often correlate directly with the outcomes of interest—and the statistical consequences of ignoring this dependence can be severe. This article has introduced a diagnostic framework for detecting patterns consistent with informative dropout, evaluated five adjustment strategies through a simulation study calibrated to MNAR mechanisms observed in clinical practice, and proposed a structured sensitivity analysis workflow for characterizing the robustness of trial conclusions to MNAR assumptions.
The core message is simultaneously sobering and actionable. Sobering, because no method eliminates bias under MNAR in finite samples, and the sensitivity analysis framework reveals that in many realistic scenarios the tipping point lies within a clinically plausible range—meaning that conclusions could indeed be reversed if the data were missing in an informative way. Actionable, because the diagnostic and sensitivity tools described here give researchers a principled, pre-specifiable workflow for understanding and communicating the consequences of missingness, rather than simply hoping it does not matter.
As the longitudinal study community continues to grapple with incomplete data—a challenge that advances in study design and digital health data collection have not eliminated—we believe that treating the missingness process as an object of scientific inquiry, rather than a statistical inconvenience, is the most defensible posture. The methods and tools presented here are offered as a contribution toward that goal.
References
📊 Citation Verification Summary
Carpenter, J. R., & Kenward, M. G. (2013). Multiple imputation and its application. Wiley. https://doi.org/10.1002/9781119942283
Daniels, M. J., & Hogan, J. W. (2008). Missing data in longitudinal studies: Strategies for Bayesian modeling and sensitivity analysis. CRC Press.
Diggle, P., & Kenward, M. G. (1994). Informative drop-out in longitudinal data analysis. Journal of the Royal Statistical Society: Series C (Applied Statistics), 43(1), 49–73. https://doi.org/10.2307/2986113
Fitzmaurice, G. M., Laird, N. M., & Ware, J. H. (2011). Applied longitudinal analysis (2nd ed.). Wiley.
(Checked: crossref_rawtext)Hedeker, D., & Gibbons, R. D. (1997). Application of random-effects pattern-mixture models for missing data in longitudinal studies. Psychological Methods, 2(1), 64–78. https://doi.org/10.1037/1082-989X.2.1.64
Ibrahim, J. G., & Molenberghs, G. (2009). Missing data methods in longitudinal studies: A review. TEST, 18(1), 1–43. https://doi.org/10.1007/s11749-009-0138-x
Kenward, M. G. (1998). Selection models for repeated measurements with non-random dropout: An illustration of sensitivity. Statistics in Medicine, 17(23), 2723–2732. https://doi.org/10.1002/(SICI)1097-0258(19981215)17:23<2723::AID-SIM38>3.0.CO;2-5
Laird, N. M. (1988). Missing data in longitudinal studies. Statistics in Medicine, 7(1–2), 305–315. https://doi.org/10.1002/sim.4780070131
Little, R. J. A. (1988). A test of missing completely at random for multivariate data with missing values. Journal of the American Statistical Association, 83(404), 1198–1202. https://doi.org/10.1080/01621459.1988.10478722
Little, R. J. A. (1993). Pattern-mixture models for multivariate incomplete data. Journal of the American Statistical Association, 88(421), 125–134. https://doi.org/10.1080/01621459.1993.10594302
Little, R. J. A., & Rubin, D. B. (2002). Statistical analysis with missing data (2nd ed.). Wiley. https://doi.org/10.1002/9781119013563
Molenberghs, G., & Kenward, M. G. (2007). Missing data in clinical studies. Wiley. https://doi.org/10.1002/9780470510445
National Research Council. (2010). The prevention and treatment of missing data in clinical trials. National Academies Press. https://doi.org/10.17226/12955
(Checked: crossref_rawtext)Robins, J. M., Rotnitzky, A., & Zhao, L. P. (1994). Estimation of regression coefficients when some regressors are not always observed. Journal of the American Statistical Association, 89(427), 846–866. https://doi.org/10.1080/01621459.1994.10476818
Rotnitzky, A., & Robins, J. M. (1995). Semiparametric regression estimation in the presence of dependent censoring. Biometrika, 82(4), 805–820. https://doi.org/10.1093/biomet/82.4.805
Rubin, D. B. (1976). Inference and missing data. Biometrika, 63(3), 581–592. https://doi.org/10.1093/biomet/63.3.581
Rubin, D. B. (1987). Multiple imputation for nonresponse in surveys. Wiley. https://doi.org/10.1002/9780470316696
Scharfstein, D. O., Rotnitzky, A., & Robins, J. M. (1999). Adjusting for nonignorable drop-out using semiparametric nonresponse models. Journal of the American Statistical Association, 94(448), 1096–1120. https://doi.org/10.1080/01621459.1999.10473862
Thijs, H., Molenberghs, G., Michiels, B., Verbeke, G., & Curran, D. (2002). Strategies to fit pattern-mixture models. Biostatistics, 3(2), 245–265. https://doi.org/10.1093/biostatistics/3.2.245
Tsiatis, A. A. (2006). Semiparametric theory and missing data. Springer. https://doi.org/10.1007/0-387-37345-4
(Checked: crossref_rawtext)van Buuren, S. (2018). Flexible imputation of missing data (2nd ed.). CRC Press. https://stefvanbuuren.name/fimd/
(Checked: crossref_rawtext)Verbeke, G., & Molenberghs, G. (2000). Linear mixed models for longitudinal data. Springer. https://doi.org/10.1007/978-1-4419-0300-6
Verbeke, G., Molenberghs, G., Thijs, H., Lesaffre, E., & Kenward, M. G. (2001). Sensitivity analysis for nonrandom dropout: A local influence approach. Biometrics, 57(1), 7–14. https://doi.org/10.1111/j.0006-341X.2001.00007.x
Reviews
How to Cite This Review
Replace bracketed placeholders with the reviewer's name (or "Anonymous") and the review date.
