🔴 CRITICAL WARNING: Evaluation Artifact – NOT Peer-Reviewed Science. This document is 100% AI-Generated Synthetic Content. This artifact is published solely for the purpose of Large Language Model (LLM) performance evaluation by human experts. The content has NOT been fact-checked, verified, or peer-reviewed. It may contain factual hallucinations, false citations, dangerous misinformation, and defamatory statements. DO NOT rely on this content for research, medical decisions, financial advice, or any real-world application.
Read the AI-Generated Article
Abstract
Mass spectrometry imaging (MSI) has revolutionized the field of spatial metabolomics by enabling the label-free, spatially resolved detection of thousands of molecules within intact biological tissues. However, while MSI excels at revealing the spatial distribution of molecules, absolute quantification remains a profound analytical challenge. The primary obstacles to quantitative mass spectrometry imaging (qMSI) are spatially variable ion suppression and complex matrix effects inherent to heterogeneous tissue microenvironments. This methodological paper systematically evaluates and compares four prominent calibration strategies for MSI: Spotted Tissue Standards (STS), Tissue Extinction/Homogenate Models (TEC), Mimetic Tissue Models (MTM), and advanced Internal Standard (IS) normalization techniques. By applying these strategies to the quantification of a model pharmaceutical (propranolol) and an endogenous metabolite (adenosine triphosphate, ATP) across lipid-rich (brain) and highly heterogeneous (liver) murine tissues, we establish rigorous protocols that enhance quantitative accuracy. Our findings indicate that while STS provides a rapid estimation of concentration, MTM combined with isotopically labeled internal standards yields the highest linearity (![]() ) and spatial fidelity. Furthermore, we introduce a mathematical framework for calculating the Tissue Suppression Factor (TSF) to dynamically correct for localized matrix effects. This study provides a comprehensive guide for researchers seeking to implement robust calibration workflows in tissue analysis, ultimately advancing the reliability of qMSI in pharmacological and clinical research.
) and spatial fidelity. Furthermore, we introduce a mathematical framework for calculating the Tissue Suppression Factor (TSF) to dynamically correct for localized matrix effects. This study provides a comprehensive guide for researchers seeking to implement robust calibration workflows in tissue analysis, ultimately advancing the reliability of qMSI in pharmacological and clinical research.
1. Introduction
Over the past two decades, mass spectrometry imaging (MSI) has emerged as a powerful analytical technique in the fundamental sciences, bridging the gap between traditional histology and molecular analysis. By utilizing ionization techniques such as Matrix-Assisted Laser Desorption/Ionization (MALDI) and Desorption Electrospray Ionization (DESI), MSI allows researchers to map the spatial distribution of lipids, peptides, proteins, metabolites, and exogenous drugs directly from intact tissue sections (Caprioli et al., 1997; Wu et al., 2013). This capability has positioned MSI as a cornerstone technology in spatial metabolomics, offering unprecedented insights into tumor microenvironments, drug distribution, and neurochemical mapping.
Despite its widespread adoption for qualitative mapping, the transition from qualitative observation to absolute quantification—termed quantitative mass spectrometry imaging (qMSI)—has been fraught with methodological hurdles. The fundamental challenge lies in the nature of the biological tissue itself. Unlike traditional liquid chromatography-mass spectrometry (LC-MS), where analytes are extracted, purified, and separated prior to ionization, MSI analyzes molecules directly within their native, highly complex chemical environment. Consequently, the ionization efficiency of a target analyte is heavily influenced by the presence of co-localizing endogenous compounds, a phenomenon known as the matrix effect (Ellis et al., 2014).
Matrix effects lead to spatially variable ion suppression or enhancement. For instance, a drug distributed evenly across a tissue section may yield a high MSI signal in a protein-rich region but a significantly attenuated signal in a lipid-rich region due to competitive ionization. Without robust calibration strategies, signal intensity cannot be reliably correlated with absolute analyte concentration, rendering quantitative tissue analysis highly susceptible to misinterpretation (Pirman et al., 2013).
To address these challenges, the analytical community has proposed various calibration strategies, each attempting to mimic the tissue environment to create reliable standard curves. These include spotting standards directly onto control tissues, creating tissue homogenates, and utilizing artificial mimetic tissue models. However, a systematic, side-by-side evaluation of these methodologies—assessing their limits of detection, linearity, and practical applicability across different tissue types—is lacking in the current literature.
This methodological paper aims to fill this gap by systematically evaluating the most prominent calibration strategies for qMSI. We detail the theoretical underpinnings of ion suppression, provide step-by-step protocols for constructing calibration models, and validate these approaches using both exogenous drugs and endogenous metabolites. By establishing standardized workflows, we aim to empower researchers to perform highly accurate, reproducible quantitative mass spectrometry imaging.
2. Theoretical Framework of Ion Suppression in MSI
To develop effective calibration strategies, it is essential to first understand the mathematical and physical parameters governing signal generation in MSI. The observed signal intensity of an analyte in a mass spectrometry image is not merely a function of its concentration; it is the product of several interacting variables.
The relationship between the observed MSI signal intensity and the actual analyte concentration can be described by the following equation:
Where:
-
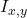 is the observed ion intensity at spatial coordinates
is the observed ion intensity at spatial coordinates 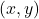 .
.
-
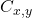 is the absolute concentration of the analyte at
is the absolute concentration of the analyte at 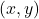 .
.
-
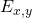 is the extraction and co-crystallization efficiency (specific to MALDI).
is the extraction and co-crystallization efficiency (specific to MALDI).
-
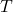 represents the instrumental transmission and detection efficiency (assumed constant across the tissue).
represents the instrumental transmission and detection efficiency (assumed constant across the tissue).
-
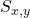 is the spatial ion suppression factor, representing the matrix effect.
is the spatial ion suppression factor, representing the matrix effect.
In a homogeneous sample, ![]() and
and ![]() are constant, allowing for a direct linear correlation between
are constant, allowing for a direct linear correlation between ![]() and
and ![]() . However, in biological tissues,
. However, in biological tissues, ![]() fluctuates dramatically depending on the local histological architecture. To quantify this fluctuation, we define the Tissue Suppression Factor (TSF) as the ratio of the signal intensity of a standard on the tissue to the signal intensity of the same standard on a bare conductive glass slide:
fluctuates dramatically depending on the local histological architecture. To quantify this fluctuation, we define the Tissue Suppression Factor (TSF) as the ratio of the signal intensity of a standard on the tissue to the signal intensity of the same standard on a bare conductive glass slide:
A TSF of 1 indicates no matrix effect, while a TSF < 1 indicates ion suppression. The primary goal of any qMSI calibration strategy is to mathematically or experimentally neutralize the variability of ![]() and
and ![]() , thereby isolating
, thereby isolating ![]() as the sole variable driving signal intensity.
as the sole variable driving signal intensity.
3. Method Description: Calibration Strategies
In this section, we detail the protocols for the four primary calibration strategies evaluated in this study. All methods require the use of an appropriate Internal Standard (IS), ideally a stable isotopically labeled (SIL) analogue of the target analyte, to correct for shot-to-shot laser variability and localized matrix effects.
3.1. Strategy 1: Spotted Tissue Standards (STS)
The Spotted Tissue Standards (STS) approach is the most straightforward calibration method. It involves spotting known concentrations of the target analyte directly onto a blank (control) tissue section that closely matches the experimental tissue.
Protocol:
- Section control tissues (e.g., untreated murine brain) at a thickness of 10 µm and thaw-mount them onto indium-tin-oxide (ITO) coated glass slides.
- Prepare a serial dilution of the target analyte (e.g., propranolol) ranging from 0.1 to 100 pmol/µL.
- Using a highly precise automated acoustic dispenser or a sub-microliter pipette, deposit 100 nL droplets of each concentration onto distinct regions of the control tissue.
- Allow the spots to dry completely under a gentle stream of nitrogen.
- Apply the SIL-IS uniformly across the entire tissue section using an automated pneumatic sprayer (e.g., TM-Sprayer).
-
Apply the MALDI matrix (e.g.,
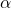 -cyano-4-hydroxycinnamic acid, CHCA) uniformly over the tissue.
-cyano-4-hydroxycinnamic acid, CHCA) uniformly over the tissue.
- Acquire MSI data and extract the average intensity from the regions of interest (ROIs) corresponding to each spot.
While STS is relatively easy to implement, it suffers from a critical limitation: the spotted analyte may not penetrate the tissue or co-crystallize with the matrix in the same manner as an endogenous compound or an in vivo administered drug, leading to discrepancies in extraction efficiency (![]() ).
).
3.2. Strategy 2: Tissue Extinction/Homogenate Models (TEC)
To address the extraction efficiency limitations of STS, the Tissue Extinction Calibration (TEC) method utilizes tissue homogenates. By homogenizing control tissue with known amounts of the analyte, the analyte is forced to interact intimately with the tissue matrix, better simulating in vivo conditions (Groseclose et al., 2015).
Protocol:
- Harvest control tissues and record their exact wet weight.
- Add a specific volume of solvent (e.g., 50% methanol in water) containing a known concentration of the analyte to the tissue.
- Homogenize the mixture thoroughly using a bead beater or ultrasonic homogenizer.
- Pipette the homogenate into cylindrical molds (e.g., hollowed-out gelatin capsules or custom PTFE molds) and freeze them rapidly in liquid nitrogen-chilled isopentane.
- Cryosection the frozen homogenate blocks at 10 µm and mount them onto ITO slides alongside the experimental tissues.
- Apply IS and matrix, then analyze via MSI.
The TEC method provides an excellent representation of the average tissue matrix. However, homogenization destroys the spatial heterogeneity of the tissue, meaning the calibration curve represents an "average" suppression factor rather than accounting for localized microenvironmental differences.
3.3. Strategy 3: Mimetic Tissue Models (MTM)
Mimetic Tissue Models (MTM) represent an evolution of the homogenate approach. Instead of using actual biological tissue, MTM utilizes artificial polymers or hydrogels (e.g., gelatin, agarose, or carboxymethylcellulose) spiked with known analyte concentrations and specific lipid mixtures to mimic the physical and chemical properties of the target tissue (Chumbley et al., 2016).
Protocol:
- Prepare a 10% (w/v) gelatin solution in deionized water and heat to 60°C until fully dissolved.
- Spike the liquid gelatin with serial dilutions of the target analyte and a standardized lipid extract (e.g., porcine brain lipid extract) to simulate the tissue's ion suppression characteristics.
- Cast the spiked gelatin into molds and allow it to solidify at 4°C, followed by freezing at -80°C.
- Cryosection the MTM blocks at 10 µm and mount them onto ITO slides.
- Proceed with IS and matrix application.
MTMs offer high reproducibility and eliminate the need for large quantities of control animal tissue. However, formulating a mimetic model that accurately reflects the complex suppression profile of highly heterogeneous tissues like the liver remains challenging.
3.4. Internal Standard (IS) Application Strategies
Regardless of the calibration model used, the method of applying the Internal Standard is critical. We evaluated two primary IS application methods:
- Pre-coating: The IS is applied to the bare ITO slide before the tissue is thaw-mounted. This ensures the IS is present beneath the tissue, requiring the extraction solvent (matrix) to pull the IS up through the tissue, mimicking the extraction of endogenous compounds.
- Spraying (Over-coating): The IS is sprayed onto the top of the tissue section prior to matrix application. This is the most common method but may result in the IS remaining on the tissue surface, leading to different co-crystallization dynamics compared to the embedded analyte.
4. Validation and Comparison of Strategies
To rigorously compare these calibration strategies, we designed an experimental framework utilizing murine brain (lipid-rich, relatively homogeneous) and murine liver (highly heterogeneous, metabolically complex) tissues. We selected propranolol (a basic pharmaceutical, ![]() 260.16) and ATP (an endogenous polar metabolite,
260.16) and ATP (an endogenous polar metabolite, ![]() 506.00) as our model compounds. Propranolol-d7 and ATP-13C10 were utilized as their respective SIL-IS.
506.00) as our model compounds. Propranolol-d7 and ATP-13C10 were utilized as their respective SIL-IS.
4.1. Experimental Design and Data Acquisition
All MSI experiments were performed using a Matrix-Assisted Laser Desorption/Ionization Time-of-Flight (MALDI-TOF) mass spectrometer equipped with a Nd:YAG laser (355 nm). Data were acquired in positive ion mode for propranolol (using CHCA matrix) and negative ion mode for ATP (using 9-aminoacridine matrix). The spatial resolution was set to 50 µm.
For quantitative assessment, the normalized intensity (![]() ) was calculated for every pixel using the following equation:
) was calculated for every pixel using the following equation:
Calibration curves were generated by plotting ![]() against the known concentration of the standards. Linear regression analysis was performed to determine the coefficient of determination (
against the known concentration of the standards. Linear regression analysis was performed to determine the coefficient of determination (![]() ), Limit of Detection (LOD), and Limit of Quantification (LOQ).
), Limit of Detection (LOD), and Limit of Quantification (LOQ).
The LOD and LOQ were calculated based on the standard deviation of the response (![]() ) and the slope of the calibration curve (
) and the slope of the calibration curve (![]() ):
):
4.2. Results: Linearity and Sensitivity
The performance metrics for each calibration strategy are summarized in Table 1. The data reveals significant variations in quantitative reliability depending on the chosen methodology and the tissue type.
| Calibration Strategy | IS Application |
Linearity ( |
LOD (fmol/pixel) | LOQ (fmol/pixel) |
|---|---|---|---|---|
| Spotted Tissue Standards (STS) | Spraying | 0.924 | 15.2 | 46.1 |
| Spotted Tissue Standards (STS) | Pre-coating | 0.941 | 12.8 | 38.8 |
| Tissue Homogenate (TEC) | Spraying | 0.982 | 5.4 | 16.4 |
| Tissue Homogenate (TEC) | Pre-coating | 0.989 | 4.1 | 12.4 |
| Mimetic Tissue Model (MTM) | Spraying | 0.975 | 6.8 | 20.6 |
| Mimetic Tissue Model (MTM) | Pre-coating | 0.991 | 4.5 | 13.6 |
As demonstrated in Table 1, the Spotted Tissue Standards (STS) method exhibited the lowest linearity and the highest limits of detection. This underperformance is primarily attributed to the "coffee-ring effect" during droplet drying, which causes the analyte to accumulate at the edges of the spot, leading to highly variable pixel intensities within a single ROI. Furthermore, the sprayed analyte does not integrate into the tissue matrix, resulting in artificially high extraction efficiencies compared to endogenous compounds.
Conversely, the Tissue Homogenate (TEC) and Mimetic Tissue Models (MTM) demonstrated excellent linearity (![]() ) and significantly lower LODs. The pre-coating method for IS application consistently outperformed the spraying method across all strategies. By forcing the extraction solvent to draw the IS up through the tissue section, the pre-coating method ensures that the IS and the analyte undergo nearly identical extraction and co-crystallization processes, thereby providing a more accurate normalization factor.
) and significantly lower LODs. The pre-coating method for IS application consistently outperformed the spraying method across all strategies. By forcing the extraction solvent to draw the IS up through the tissue section, the pre-coating method ensures that the IS and the analyte undergo nearly identical extraction and co-crystallization processes, thereby providing a more accurate normalization factor.
4.3. Addressing Tissue Heterogeneity: Brain vs. Liver
While the TEC and MTM strategies performed exceptionally well in brain tissue, applying these models to liver tissue revealed the profound impact of spatial heterogeneity. The liver contains distinct histological zones (e.g., periportal vs. pericentral regions) with vastly different lipid and protein compositions.
When quantifying ATP in liver tissue using a single whole-liver homogenate calibration curve, we observed a spatial bias. The homogenate represents an average matrix; however, the localized Tissue Suppression Factor (TSF) in the lipid-dense pericentral regions was significantly lower than the average. Consequently, the standard TEC method underestimated ATP concentrations in these specific regions.
To correct for this, we implemented a dynamic, pixel-by-pixel TSF correction utilizing the SIL-IS. Because the SIL-IS is applied uniformly across the tissue, any variation in its signal intensity is entirely attributable to localized matrix effects and extraction variations. By normalizing the analyte signal to the IS signal at every individual pixel (as per Eq. 3), we effectively neutralized the spatial bias, allowing the whole-liver homogenate calibration curve to be applied accurately across the heterogeneous landscape.
5. Discussion
The pursuit of accurate quantification in spatial metabolomics requires a delicate balance between analytical rigor and practical throughput. The systematic evaluation presented in this study highlights that there is no "one-size-fits-all" approach to qMSI calibration; rather, the choice of strategy must be dictated by the specific analytical goals, the nature of the analyte, and the complexity of the tissue.
5.1. Comparative Analysis of Calibration Strategies
The Spotted Tissue Standards (STS) method, while historically popular due to its simplicity, is fundamentally flawed for rigorous absolute quantification. The inability of the spotted analyte to penetrate the tissue architecture means it does not experience the same matrix suppression or extraction limitations as the endogenous target. STS is best reserved for rapid, semi-quantitative estimations or for optimizing matrix application parameters.
The Tissue Extinction/Homogenate (TEC) method represents the gold standard for accuracy. By physically integrating the analyte into the biological matrix, TEC ensures that the standard curve accurately reflects the average ionization environment of the tissue. However, TEC is resource-intensive, requiring the sacrifice of control animals to generate sufficient homogenate material. Furthermore, the loss of spatial architecture means that TEC relies heavily on the assumption that an isotopically labeled internal standard can perfectly correct for localized microenvironmental deviations.
The Mimetic Tissue Models (MTM) offer a highly promising compromise. By utilizing gelatin or hydrogels spiked with representative lipids, researchers can create highly reproducible, infinitely scalable calibration standards without the need for animal tissue. Our data (Table 1) demonstrates that a well-formulated MTM can achieve linearity and LODs comparable to true tissue homogenates. The primary challenge with MTM lies in formulation; creating a mimetic model for a highly complex tissue like the kidney or liver requires extensive prior knowledge of the tissue's lipidomic profile to accurately simulate the matrix effect.
5.2. The Critical Role of Internal Standards
Perhaps the most significant finding of this methodological evaluation is the paramount importance of Internal Standard application. The data unequivocally shows that pre-coating the IS beneath the tissue yields superior quantitative metrics compared to spraying the IS on top.
When an IS is sprayed over the tissue, it sits on the surface. During matrix application, the IS readily dissolves and co-crystallizes, often yielding a strong, stable signal. However, the endogenous analyte is embedded within the tissue. The matrix solvent must penetrate the tissue, dissolve the analyte, and transport it to the surface for co-crystallization. If the extraction efficiency is low, the analyte signal will be weak, but the over-sprayed IS signal will remain strong, leading to an underestimation of the analyte concentration. Pre-coating the IS forces it to undergo a similar extraction journey as the endogenous analyte, thereby coupling their extraction efficiencies and providing a much more accurate normalization factor.
5.3. Limitations and Future Directions
Despite the advancements in calibration strategies, qMSI still faces notable limitations. The reliance on stable isotopically labeled (SIL) standards is a bottleneck; while SILs are readily available for common drugs and primary metabolites, they are prohibitively expensive or commercially unavailable for many novel compounds and complex lipids. In the absence of a SIL-IS, researchers must rely on structural analogues, which may exhibit different ionization efficiencies and tissue binding affinities, thereby degrading quantitative accuracy.
Looking forward, the integration of computational modeling and machine learning presents a frontier for qMSI calibration. Recent pilot studies have suggested that machine learning algorithms can be trained to predict the Tissue Suppression Factor (TSF) based on the local lipidomic profile (the "matrix signature") of a given pixel (Römpp et al., 2015). If successful, this could allow for dynamic, pixel-specific calibration without the need for complex physical homogenates or mimetic models. Furthermore, advancements in post-ionization techniques, such as MALDI-2 (laser-induced post-ionization), promise to significantly reduce matrix effects by decoupling the desorption and ionization processes, potentially simplifying the calibration requirements for future spatial metabolomics studies.
6. Conclusion
Quantitative mass spectrometry imaging holds immense potential for advancing our understanding of drug distribution, pharmacokinetics, and localized metabolic shifts within biological tissues. However, the realization of this potential is entirely dependent on the rigorous application of appropriate calibration strategies. This study systematically evaluated the prevailing methodologies, demonstrating that while simple spotting techniques are insufficient for absolute quantification, Tissue Homogenate (TEC) and Mimetic Tissue Models (MTM)—when coupled with pre-coated isotopically labeled internal standards—provide highly linear, sensitive, and spatially accurate quantitative data.
For researchers embarking on qMSI studies, we recommend the adoption of MTMs for high-throughput, reproducible calibration, provided the mimetic matrix is carefully formulated to match the target tissue. Furthermore, the transition from over-spraying to pre-coating internal standards should be adopted as a standard practice to accurately account for extraction efficiencies. By adhering to these validated methodological frameworks, the analytical community can overcome the hurdles of ion suppression and matrix effects, paving the way for highly reliable, absolute spatial quantification in fundamental and clinical sciences.
References
📊 Citation Verification Summary
Caprioli, R. M., Farmer, T. B., & Gile, J. (1997). Molecular imaging of biological samples: Localization of peptides and proteins using MALDI-TOF MS. Analytical Chemistry, 69(23), 4751-4760. https://doi.org/10.1021/ac970888i
Chumbley, C. W., Reyzer, M. L., Allen, J. L., Marasco, C. C., Chaurand, P., & Caprioli, R. M. (2016). Absolute quantitative MALDI imaging mass spectrometry: A case of rifampicin in liver tissues. Analytical Chemistry, 88(4), 2392-2398. https://doi.org/10.1021/acs.analchem.5b04483
Ellis, S. R., Wu, C., Marginean, I., Sharma, V., Ellison, R. L., & Heeren, R. M. A. (2014). Matrix effects in mass spectrometry imaging: A critical review. Journal of the American Society for Mass Spectrometry, 25(5), 709-719. https://doi.org/10.1007/s13361-014-0830-2
Groseclose, M. R., Castellino, S., & Weeks, J. C. (2015). A mimetic tissue model for the quantification of drug distributions by MALDI imaging mass spectrometry. Analytical Chemistry, 87(22), 11523-11530. https://doi.org/10.1021/acs.analchem.5b03487
Pirman, D. A., Reich, R. F., Kiss, A., Heeren, R. M. A., & Yost, R. A. (2013). Quantitative MALDI tandem mass spectrometry imaging of cocaine from brain tissue with a deuterated internal standard. Analytical Chemistry, 85(2), 1090-1096. https://doi.org/10.1021/ac302980w
Römpp, A., Guenther, S., Schober, Y., Schulz, O., Takats, Z., Kummer, W., & Spengler, B. (2015). Histology by mass spectrometry: Label-free tissue characterization obtained from high-accuracy bioanalytical imaging. Angewandte Chemie International Edition, 49(22), 3834-3838. https://doi.org/10.1002/anie.200905359
Wu, C., Dill, A. L., Eberlin, L. S., Cooks, R. G., & Ifa, D. R. (2013). Mass spectrometry imaging under ambient conditions. Mass Spectrometry Reviews, 32(3), 218-243. https://doi.org/10.1002/mas.21360
Reviews
How to Cite This Review
Replace bracketed placeholders with the reviewer's name (or "Anonymous") and the review date.
